When people think about scientific research, they often imagine complex equipment, long experiments, and data that only scientists understand. But behind every lab experiment lies a bigger purpose: improving real lives.
As a pharmacy student at RCSI University of Medicine and Health Sciences, I experienced this connection firsthand. During my studies, I carried out research in the RCSI Cancer Bioengineering Lab as part of my thesis work. This gave me the opportunity to see how laboratory science can contribute to real advances in healthcare.
Starting in the Lab
My project focused on developing new materials that could support therapeutic cancer vaccines (adjuvants). These vaccines are different from traditional ones because, instead of preventing disease, they are designed to help the body fight cancer more effectively.
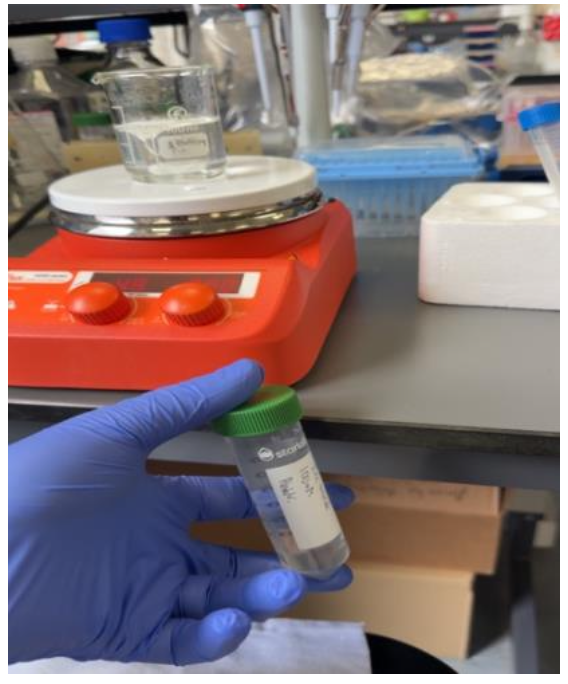
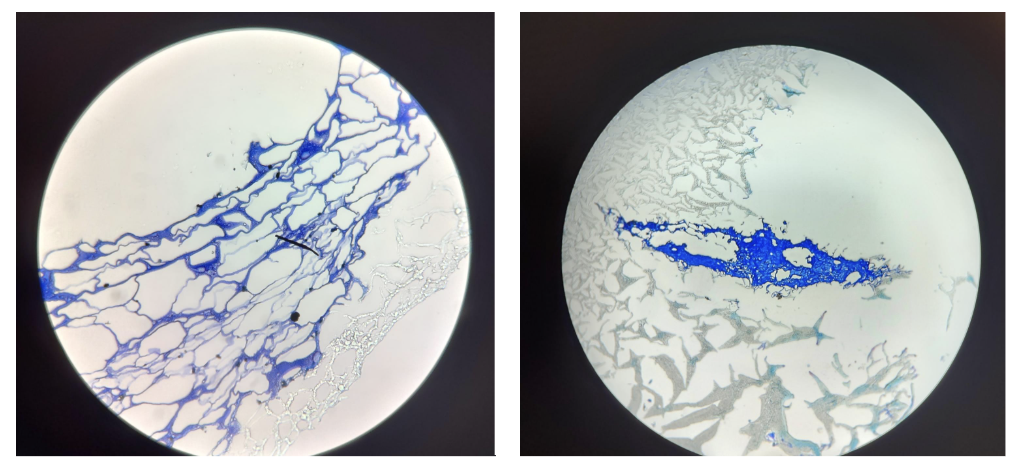
At the centre of this research is alginate, a natural material derived from brown algae. Although simple in origin, it can be modified into tiny gel-like particles with immune-stimulating potential.
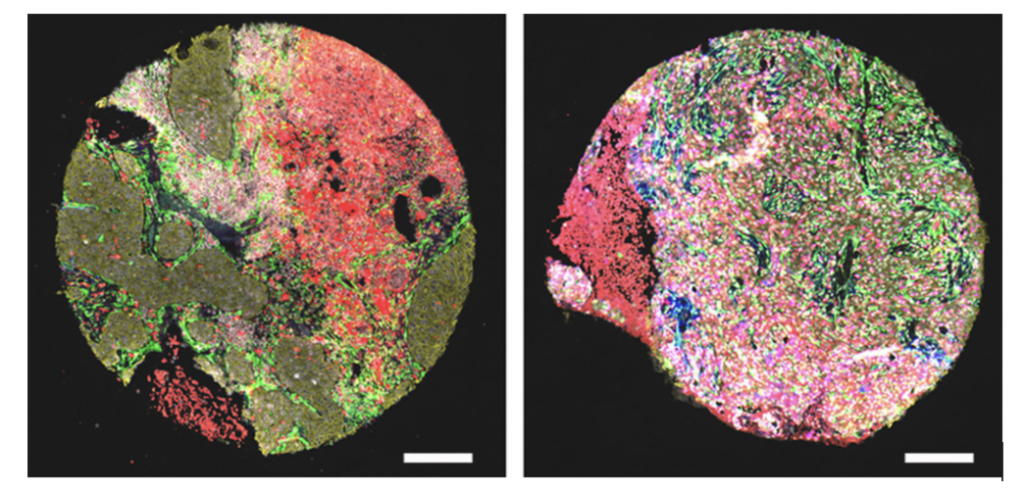
In the lab, this involved preparing solutions, carefully mixing components, and analysing the particles using techniques such as DLS and NTA. Small changes in concentration or preparation could completely alter the outcome, which required patience, precision, and problem-solving.
More Than Just Experiments
At first, lab work can feel like simply following steps: weighing chemicals, stirring solutions, and running instruments. But over time, I began to understand the bigger picture. This is where research becomes meaningful — each small experiment contributes to the larger goal of developing life-changing therapeutics.
Challenges Along the Way
Research is not always straightforward. Not every experiment works as expected. Sometimes results are unclear, and methods need to be adjusted and repeated.
During my project, I had to optimise various conditions to achieve stable, effective particles while also reviewing previous studies in this field. This involved testing multiple concentrations and analysing their behaviour. It taught me that failure is not a setback, but an essential part of discovery.
Connecting to Real Life
The most important lesson I learned is that research is never just about the lab.
The materials we develop could one day be used to enhance cancer treatments, improving patient outcomes and quality of life. While my work is still in its early stages, it contributes to a broader effort to make therapies more effective and targeted.
As a future pharmacist, this connection is especially important. It highlights how science, healthcare, and patient care are all deeply linked.
From Student to Scientist
This experience has changed how I view research. It is no longer just something I study — it is something I actively contribute to.
It has also strengthened my interest in pharmaceutical research, showing me how lab innovation can eventually translate into real treatments for patients.
Final Thoughts
Research may begin in the lab, but its impact reaches far beyond it.
Every measurement, every experiment, and every challenge faced is part of a journey toward improving health and saving lives. For students like me, it is a reminder that what we do today in the lab could shape the treatments of tomorrow.
Written by Abdulrahman Alowaihan